Metagenomic all-vs-all sample comparison, 10x faster
K-mers, subsequences of length k, are a fundamental building block of genomic data analysis with various applications. One such application is measuring the distance between different metagenomic NGS samples, for example to perform clustering. We can simply represent each sample as a vector of counted k-mers – a k-mer spectrum – for some k, and compute the distance between different vectors. However, this raises two problems. How efficiently can we compute such distances? The vectors might contain billions of entries, and all-vs-all distance is a problem that scales quadratically in the number of samples. And which distance metric is most accurate from a biological perspective - Euclidean, cosine, Jaccard, or some other distance metric?
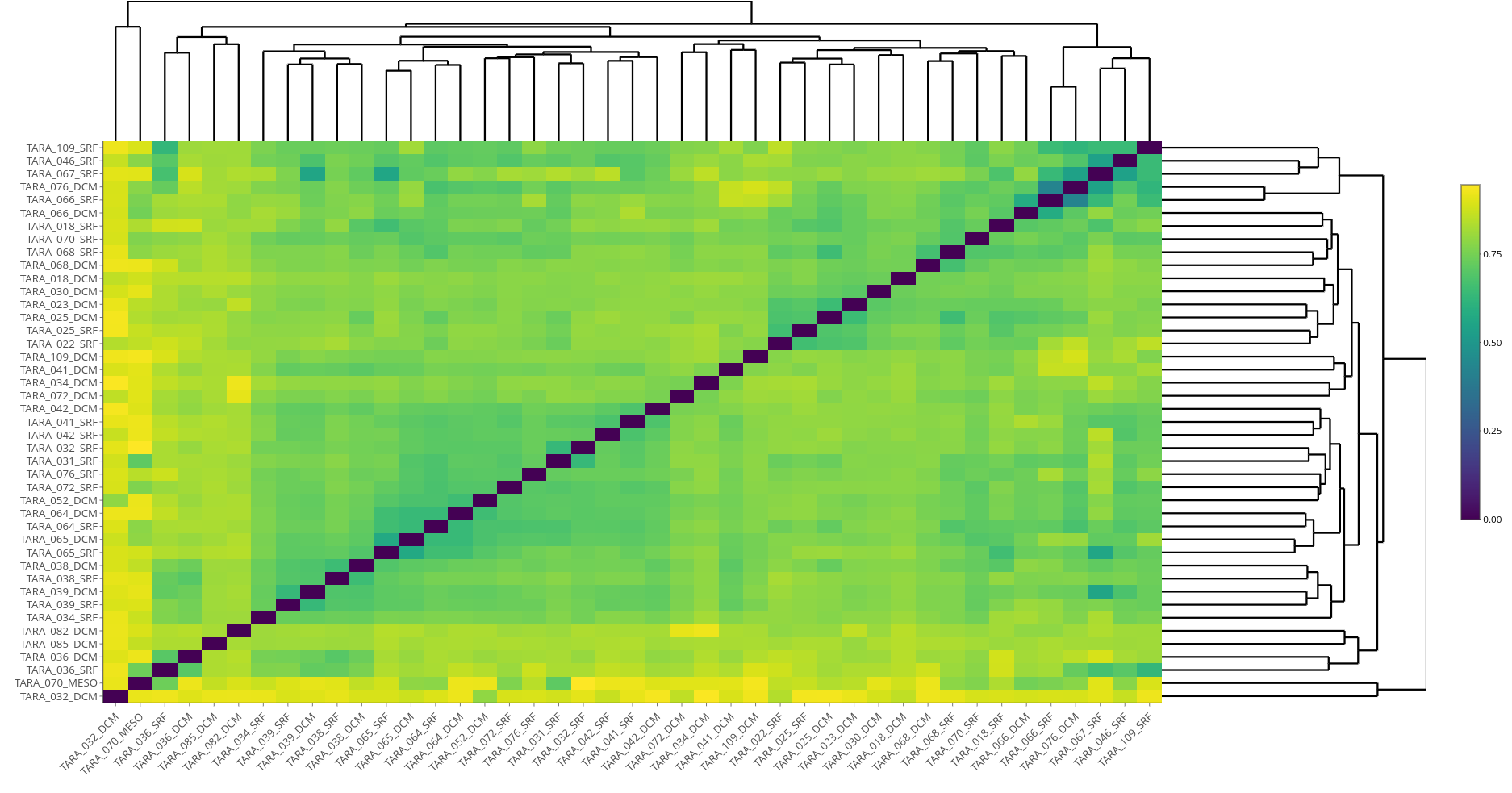
In their impressive paper Libra: scalable k-mer based tool for massive all-vs-all metagenome comparisons (GigaScience 2019), Choi et al implement a k-mer based metagenomic distance analysis tool on Apache Hadoop. They demonstrate that on a large dataset, the Tara Oceans viral metagenome (TOV), for 43 samples, the all-vs-all distance matrix can be computed efficiently. They also use various kinds of data to experimentally determine the best distance metric, and find that cosine distance with logarithmic weighting is the most biologically accurate way to compare metagenomes.
At JNP Solutions, we develop scalable k-mer analysis tools for Apache Spark. Recently, as a new test case, we implemented the all-against-all distance calculation for k-mer vectors in our tool Hypercut to compare our results against Libra.
Hypercut offers superior speed and compression. Based on the authors’ reported runtime, in Libra, library construction took 1782 CPU hours for the 43 TOV samples. Hypercut can construct the same library (vectors of counted k-mers) in 128 CPU hours (1.1 TB of input data). In Libra, all vs all distance computation took 162 CPU hours; in Hypercut it takes 55 CPU hours. So the total time taken for indexing and distance computation is 1944h in Libra vs 183h in Hypercut: more than a 10x improvement.
The disk space used by Libra’s index is 1124 GB, whereas Hypercut’s is 144 GB. These differences in performance are a result of our ongoing intensive research and algorithm development.
| Libra | Hypercut | |
|---|---|---|
| Index construction (CPUh) | 1782 | 128 |
| All-vs-all comparison (CPUh) | 162 | 55 |
| Total time (CPUh) | 1944 | 183 |
| Disk space for index (GB) | 1124 | 144 |
Hypercut runs efficiently on all major cloud platforms (or anywhere that Spark will run, including on normal PCs and clusters). We are pleased to be able to offer it to our customers today. For more information, or to schedule a demo, please contact us at info@jnpsolutions.io.